Boost Your Data Munging With R
As a language, R is strongly tied to data and is thus used mostly by statisticians and data scientists. Many who already use R for machine learning, though, are not aware that data munging can be done faster in R, meaning another tool is not required for that task.
In this article, Freelance Software Engineer Jan Gorecki explores tabular data transformations and introduces us to one of the fastest open-source data wrangling tools available.
As a language, R is strongly tied to data and is thus used mostly by statisticians and data scientists. Many who already use R for machine learning, though, are not aware that data munging can be done faster in R, meaning another tool is not required for that task.
In this article, Freelance Software Engineer Jan Gorecki explores tabular data transformations and introduces us to one of the fastest open-source data wrangling tools available.
Jan is a business intelligence and data warehousing expert with advanced R skills and some infrastructure experience.
Expertise
The R language is often perceived as a language for statisticians and data scientists. Quite a long time ago, this was mostly true. However, over the years the flexibility R provides via packages has made R into a more general purpose language. R was open sourced in 1995, and since that time repositories of R packages are constantly growing. Still, compared to languages like Python, R is strongly based around the data.
Speaking about data, tabular data deserves particular attention, as it’s one of the most commonly used data types. It is a data type which corresponds to a table structure known in databases, where each column can be of a different type, and processing performance of that particular data type is the crucial factor for many applications.

In this article, we are going to present how to achieve tabular data transformation in an efficient manner. Many people who use R already for machine learning are not aware that data munging can be done faster in R, and that they do not need to use another tool for it.
High-performance Solution in R
Base R introduced the data.frame class in the year 1997, which was based on S-PLUS before it. Unlike commonly used databases which store data row by row, R data.frame stores the data in memory as a column-oriented structure, thus making it more cache-efficient for column operations which are common in analytics. Additionally, even though R is a functional programming language, it does not enforce that on the developer. Both opportunities have been well addressed by data.table R package, which is available in CRAN repository. It performs quite fast when grouping operations, and is particularly memory efficient by being careful about materializing intermediate data subsets, such as materializing only those columns necessary for a certain task. It also avoids unnecessary copies through its reference semantics while adding or updating columns. The first version of the package has been published in April 2006, significantly improving data.frame performance at that time. The initial package description was:
This package does very little. The only reason for its existence is that the white book specifies that data.frame must have rownames. This package defines a new class data.table which operates just like a data.frame, but uses up to 10 times less memory, and can be up to 10 times faster to create (and copy). It also takes the opportunity to allow subset() and with() like expressions inside the []. Most of the code is copied from base functions with the code manipulating row.names removed.
Since then, both data.frame and data.table implementations have been improved, but data.table remains to be incredibly faster than base R. In fact, data.table isn’t just faster than base R, but it appears to be one of the fastest open-source data wrangling tool available, competing with tools like Python Pandas, and columnar storage databases or big data apps like Spark. Its performance over distributed shared infrastructure hasn’t been yet benchmarked, but being able to have up to two billion rows on a single instance gives promising prospects. Outstanding performance goes hand-in-hand with the functionalities. Additionally, with recent efforts at parallelizing time-consuming parts for incremental performance gains, one direction towards pushing the performance limit seems quite clear.
Data Transformation Examples
Learning R gets a little bit easier because of the fact that it works interactively, so we can follow examples step by step and look at the results of each step at any time. Before we start, let’s install the data.table package from CRAN repository.
install.packages("data.table")
Useful hint: We can open the manual of any function just by typing its name with leading question mark, i.e. ?install.packages.
Loading Data into R
There are tons of packages for extracting data from a wide range of formats and databases, which often includes native drivers. We will load data from the CSV file, the most common format for raw tabular data. File used in the following examples can be found here. We don’t have to bother about CSV reading performance as the fread function is highly optimized on that.
In order to use any function from a package, we need to load it with the library call.
library(data.table)
DT <- fread("flights14.csv")
print(DT)
## year month day dep_delay arr_delay carrier origin dest air_time
## 1: 2014 1 1 14 13 AA JFK LAX 359
## 2: 2014 1 1 -3 13 AA JFK LAX 363
## 3: 2014 1 1 2 9 AA JFK LAX 351
## 4: 2014 1 1 -8 -26 AA LGA PBI 157
## 5: 2014 1 1 2 1 AA JFK LAX 350
## ---
## 253312: 2014 10 31 1 -30 UA LGA IAH 201
## 253313: 2014 10 31 -5 -14 UA EWR IAH 189
## 253314: 2014 10 31 -8 16 MQ LGA RDU 83
## 253315: 2014 10 31 -4 15 MQ LGA DTW 75
## 253316: 2014 10 31 -5 1 MQ LGA SDF 110
## distance hour
## 1: 2475 9
## 2: 2475 11
## 3: 2475 19
## 4: 1035 7
## 5: 2475 13
## ---
## 253312: 1416 14
## 253313: 1400 8
## 253314: 431 11
## 253315: 502 11
## 253316: 659 8
If our data is not well modeled for further processing, as they need to be reshaped from long-to-wide or wide-to-long (also known as pivot and unpivot) format, we may look at ?dcast and ?melt functions, known from reshape2 package. However, data.table implements faster and memory efficient methods for data.table/data.frame class.
Querying with data.table Syntax
If You’re Familiar with data.frame
Query data.table is very similar to query data.frame. While filtering in i argument, we can use column names directly without the need to access them with the $ sign, like df[df$col > 1, ]. When providing the next argument j, we provide an expression to be evaluated in the scope of our data.table. To pass a non-expression j argument use with=FALSE. Third argument, not present in data.frame method, defines the groups, making the expression in j to be evaluated by groups.
# data.frame
DF[DF$col1 > 1L, c("col2", "col3")]
# data.table
DT[col1 > 1L, .(col2, col3), ...] # by group using: `by = col4`
If You’re Familiar with Databases
Query data.table in many aspects corresponds to SQL queries that more people might be familiar with. DT below represents data.table object and corresponds to SQLs FROM clause.
DT[ i = where,
j = select | update,
by = group by]
[ having, ... ]
[ order by, ... ]
[ ... ] ... [ ... ]

Sorting Rows and Re-Ordering Columns
Sorting data is a crucial transformation for time series, and it is also imports for data extract and presentation. Sort can be achieved by providing the integer vector of row order to i argument, the same way as data.frame. First argument in query order(carrier, -dep_delay) will select data in ascending order on carrier field and descending order on dep_delay measure. Second argument j, as described in the previous section, defines the columns (or expressions) to be returned and their order.
ans <- DT[order(carrier, -dep_delay),
.(carrier, origin, dest, dep_delay)]
head(ans)
## carrier origin dest dep_delay
## 1: AA EWR DFW 1498
## 2: AA JFK BOS 1241
## 3: AA EWR DFW 1071
## 4: AA EWR DFW 1056
## 5: AA EWR DFW 1022
## 6: AA EWR DFW 989
To re-order data by reference, instead of querying data in specific order, we use set* functions.
setorder(DT, carrier, -dep_delay)
leading.cols <- c("carrier","dep_delay")
setcolorder(DT, c(leading.cols, setdiff(names(DT), leading.cols)))
print(DT)
## carrier dep_delay year month day arr_delay origin dest air_time
## 1: AA 1498 2014 10 4 1494 EWR DFW 200
## 2: AA 1241 2014 4 15 1223 JFK BOS 39
## 3: AA 1071 2014 6 13 1064 EWR DFW 175
## 4: AA 1056 2014 9 12 1115 EWR DFW 198
## 5: AA 1022 2014 6 16 1073 EWR DFW 178
## ---
## 253312: WN -12 2014 3 9 -21 LGA BNA 115
## 253313: WN -13 2014 3 10 -18 EWR MDW 112
## 253314: WN -13 2014 5 17 -30 LGA HOU 202
## 253315: WN -13 2014 6 15 10 LGA MKE 101
## 253316: WN -13 2014 8 19 -30 LGA CAK 63
## distance hour
## 1: 1372 7
## 2: 187 13
## 3: 1372 10
## 4: 1372 6
## 5: 1372 7
## ---
## 253312: 764 16
## 253313: 711 20
## 253314: 1428 17
## 253315: 738 20
## 253316: 397 16
Most often, we don’t need both the original dataset and the ordered/sorted dataset. By default, the R language, similar to other functional programming languages, will return sorted data as new object, and thus will require twice as much memory as sorting by reference.
Subset Queries
Let’s create a subset dataset for flight origin “JFK” and month from 6 to 9. In the second argument, we subset results to listed columns, adding one calculated variable sum_delay.
ans <- DT[origin == "JFK" & month %in% 6:9,
.(origin, month, arr_delay, dep_delay, sum_delay = arr_delay + dep_delay)]
head(ans)
## origin month arr_delay dep_delay sum_delay
## 1: JFK 7 925 926 1851
## 2: JFK 8 727 772 1499
## 3: JFK 6 466 451 917
## 4: JFK 7 414 450 864
## 5: JFK 6 411 442 853
## 6: JFK 6 333 343 676
By default, when subsetting dataset on single column data.table will automatically create an index for that column. This results in real-time answers on any further filtering calls on that column.
Update Dataset
Adding a new column by reference is performed using the := operator, it assigns a variable into dataset in place. This avoids in-memory copy of dataset, so we don’t need to assign results to each new variable.
DT[, sum_delay := arr_delay + dep_delay]
head(DT)
## carrier dep_delay year month day arr_delay origin dest air_time
## 1: AA 1498 2014 10 4 1494 EWR DFW 200
## 2: AA 1241 2014 4 15 1223 JFK BOS 39
## 3: AA 1071 2014 6 13 1064 EWR DFW 175
## 4: AA 1056 2014 9 12 1115 EWR DFW 198
## 5: AA 1022 2014 6 16 1073 EWR DFW 178
## 6: AA 989 2014 6 11 991 EWR DFW 194
## distance hour sum_delay
## 1: 1372 7 2992
## 2: 187 13 2464
## 3: 1372 10 2135
## 4: 1372 6 2171
## 5: 1372 7 2095
## 6: 1372 11 1980
To add more variables at once, we can use DT[, :=(sum_delay = arr_delay + dep_delay)] syntax, similar to .(sum_delay = arr_delay + dep_delay) when querying from dataset.
It is possible to sub-assign by reference, updating only particular rows in place, just by combining with i argument.
DT[origin=="JFK",
distance := NA]
head(DT)
## carrier dep_delay year month day arr_delay origin dest air_time
## 1: AA 1498 2014 10 4 1494 EWR DFW 200
## 2: AA 1241 2014 4 15 1223 JFK BOS 39
## 3: AA 1071 2014 6 13 1064 EWR DFW 175
## 4: AA 1056 2014 9 12 1115 EWR DFW 198
## 5: AA 1022 2014 6 16 1073 EWR DFW 178
## 6: AA 989 2014 6 11 991 EWR DFW 194
## distance hour sum_delay
## 1: 1372 7 2992
## 2: NA 13 2464
## 3: 1372 10 2135
## 4: 1372 6 2171
## 5: 1372 7 2095
## 6: 1372 11 1980
Aggregate Data
To aggregate data, we provide the third argument by to the square bracket. Then, in j we need to provide aggregate function calls, so the data can be actually aggregated. The .N symbol used in the j argument corresponds to the number of all observations in each group. As previously mentioned, aggregates can be combined with subsets on rows and selecting columns.
ans <- DT[,
.(m_arr_delay = mean(arr_delay),
m_dep_delay = mean(dep_delay),
count = .N),
.(carrier, month)]
head(ans)
## carrier month m_arr_delay m_dep_delay count
## 1: AA 10 5.541959 7.591497 2705
## 2: AA 4 1.903324 3.987008 2617
## 3: AA 6 8.690067 11.476475 2678
## 4: AA 9 -1.235160 3.307078 2628
## 5: AA 8 4.027474 8.914054 2839
## 6: AA 7 9.159886 11.665953 2802
Often, we may need to compare a value of a row to its aggregate over a group. In SQL, we apply aggregates over partition by: AVG(arr_delay) OVER (PARTITION BY carrier, month).
ans <- DT[,
.(arr_delay, carrierm_mean_arr = mean(arr_delay),
dep_delay, carrierm_mean_dep = mean(dep_delay)),
.(carrier, month)]
head(ans)
## carrier month arr_delay carrierm_mean_arr dep_delay carrierm_mean_dep
## 1: AA 10 1494 5.541959 1498 7.591497
## 2: AA 10 840 5.541959 848 7.591497
## 3: AA 10 317 5.541959 338 7.591497
## 4: AA 10 292 5.541959 331 7.591497
## 5: AA 10 322 5.541959 304 7.591497
## 6: AA 10 306 5.541959 299 7.591497
If we don’t want to query data with those aggregates, and instead just put them into actual table updating by reference, we can accomplish that with := operator. This avoids the in-memory copy of the dataset, so we don’t need to assign results to the new variable.
DT[,
`:=`(carrierm_mean_arr = mean(arr_delay),
carrierm_mean_dep = mean(dep_delay)),
.(carrier, month)]
head(DT)
## carrier dep_delay year month day arr_delay origin dest air_time
## 1: AA 1498 2014 10 4 1494 EWR DFW 200
## 2: AA 1241 2014 4 15 1223 JFK BOS 39
## 3: AA 1071 2014 6 13 1064 EWR DFW 175
## 4: AA 1056 2014 9 12 1115 EWR DFW 198
## 5: AA 1022 2014 6 16 1073 EWR DFW 178
## 6: AA 989 2014 6 11 991 EWR DFW 194
## distance hour sum_delay carrierm_mean_arr carrierm_mean_dep
## 1: 1372 7 2992 5.541959 7.591497
## 2: NA 13 2464 1.903324 3.987008
## 3: 1372 10 2135 8.690067 11.476475
## 4: 1372 6 2171 -1.235160 3.307078
## 5: 1372 7 2095 8.690067 11.476475
## 6: 1372 11 1980 8.690067 11.476475
Join Datasets
Base R joining and merging of datasets is considered a special type of subset operation. We provide a dataset to which we want to join in the first square bracket argument i. For each row in dataset provided to i, we match rows from the dataset in which we use [. If we want to keep only matching rows (inner join), then we pass an extra argument nomatch = 0L. We use on argument to specify columns on which we want to join both datasets.
# create reference subset
carrierdest <- DT[, .(count=.N), .(carrier, dest) # count by carrier and dest
][1:10 # just 10 first groups
] # chaining `[...][...]` as subqueries
print(carrierdest)
## carrier dest count
## 1: AA DFW 5877
## 2: AA BOS 1173
## 3: AA ORD 4798
## 4: AA SEA 298
## 5: AA EGE 85
## 6: AA LAX 3449
## 7: AA MIA 6058
## 8: AA SFO 1312
## 9: AA AUS 297
## 10: AA DCA 172
# outer join
ans <- carrierdest[DT, on = c("carrier","dest")]
print(ans)
## carrier dest count dep_delay year month day arr_delay origin
## 1: AA DFW 5877 1498 2014 10 4 1494 EWR
## 2: AA BOS 1173 1241 2014 4 15 1223 JFK
## 3: AA DFW 5877 1071 2014 6 13 1064 EWR
## 4: AA DFW 5877 1056 2014 9 12 1115 EWR
## 5: AA DFW 5877 1022 2014 6 16 1073 EWR
## ---
## 253312: WN BNA NA -12 2014 3 9 -21 LGA
## 253313: WN MDW NA -13 2014 3 10 -18 EWR
## 253314: WN HOU NA -13 2014 5 17 -30 LGA
## 253315: WN MKE NA -13 2014 6 15 10 LGA
## 253316: WN CAK NA -13 2014 8 19 -30 LGA
## air_time distance hour sum_delay carrierm_mean_arr
## 1: 200 1372 7 2992 5.541959
## 2: 39 NA 13 2464 1.903324
## 3: 175 1372 10 2135 8.690067
## 4: 198 1372 6 2171 -1.235160
## 5: 178 1372 7 2095 8.690067
## ---
## 253312: 115 764 16 -33 6.921642
## 253313: 112 711 20 -31 6.921642
## 253314: 202 1428 17 -43 22.875845
## 253315: 101 738 20 -3 14.888889
## 253316: 63 397 16 -43 7.219670
## carrierm_mean_dep
## 1: 7.591497
## 2: 3.987008
## 3: 11.476475
## 4: 3.307078
## 5: 11.476475
## ---
## 253312: 11.295709
## 253313: 11.295709
## 253314: 30.546453
## 253315: 24.217560
## 253316: 17.038047
# inner join
ans <- DT[carrierdest, # for each row in carrierdest
nomatch = 0L, # return only matching rows from both tables
on = c("carrier","dest")] # joining on columns carrier and dest
print(ans)
## carrier dep_delay year month day arr_delay origin dest air_time
## 1: AA 1498 2014 10 4 1494 EWR DFW 200
## 2: AA 1071 2014 6 13 1064 EWR DFW 175
## 3: AA 1056 2014 9 12 1115 EWR DFW 198
## 4: AA 1022 2014 6 16 1073 EWR DFW 178
## 5: AA 989 2014 6 11 991 EWR DFW 194
## ---
## 23515: AA -8 2014 10 11 -13 JFK DCA 53
## 23516: AA -9 2014 5 21 -12 JFK DCA 52
## 23517: AA -9 2014 6 5 -6 JFK DCA 53
## 23518: AA -9 2014 10 2 -21 JFK DCA 51
## 23519: AA -11 2014 5 27 10 JFK DCA 55
## distance hour sum_delay carrierm_mean_arr carrierm_mean_dep count
## 1: 1372 7 2992 5.541959 7.591497 5877
## 2: 1372 10 2135 8.690067 11.476475 5877
## 3: 1372 6 2171 -1.235160 3.307078 5877
## 4: 1372 7 2095 8.690067 11.476475 5877
## 5: 1372 11 1980 8.690067 11.476475 5877
## ---
## 23515: NA 15 -21 5.541959 7.591497 172
## 23516: NA 15 -21 4.150172 8.733665 172
## 23517: NA 15 -15 8.690067 11.476475 172
## 23518: NA 15 -30 5.541959 7.591497 172
## 23519: NA 15 -1 4.150172 8.733665 172
Be aware that because of the consistency to base R subsetting, the outer join is by default RIGHT OUTER. If we are looking for LEFT OUTER, we need to swap the tables, as in the example above. Exact behavior can also be easily controlled in merge data.table method, using the same API as base R merge data.frame.
If we want to simply lookup the column(s) to our dataset, we can efficiently do it with := operator in j argument while joining. The same way as we sub-assign by reference, as described in the Update dataset section, we just now add a column by reference from the dataset to which we join. This avoids the in-memory copy of data, so we don’t need to assign results into new variables.
DT[carrierdest, # data.table to join with
lkp.count := count, # lookup `count` column from `carrierdest`
on = c("carrier","dest")] # join by columns
head(DT)
## carrier dep_delay year month day arr_delay origin dest air_time
## 1: AA 1498 2014 10 4 1494 EWR DFW 200
## 2: AA 1241 2014 4 15 1223 JFK BOS 39
## 3: AA 1071 2014 6 13 1064 EWR DFW 175
## 4: AA 1056 2014 9 12 1115 EWR DFW 198
## 5: AA 1022 2014 6 16 1073 EWR DFW 178
## 6: AA 989 2014 6 11 991 EWR DFW 194
## distance hour sum_delay carrierm_mean_arr carrierm_mean_dep lkp.count
## 1: 1372 7 2992 5.541959 7.591497 5877
## 2: NA 13 2464 1.903324 3.987008 1173
## 3: 1372 10 2135 8.690067 11.476475 5877
## 4: 1372 6 2171 -1.235160 3.307078 5877
## 5: 1372 7 2095 8.690067 11.476475 5877
## 6: 1372 11 1980 8.690067 11.476475 5877
For aggregate while join, use by = .EACHI. It performs join that won’t materialize intermediate join results and will apply aggregates on the fly, making it memory efficient.
Rolling join is an uncommon feature, designed for dealing with ordered data. It fits perfectly for processing temporal data, and time series in general. It basically roll matches in join condition to next matching value. Use it by providing the roll argument when joining.
Fast overlap join joins datasets based on periods and its overlapping handling by using various overlaping operators: any, within, start, end.
A non-equi join feature to join datasets using non-equal condition is currently being developed.
Profiling Data
When exploring our dataset, we may sometimes want to collect technical information on the subject, to better understand the quality of the data.
Descriptive Statistics
summary(DT)
## carrier dep_delay year month
## Length:253316 Min. :-112.00 Min. :2014 Min. : 1.000
## Class :character 1st Qu.: -5.00 1st Qu.:2014 1st Qu.: 3.000
## Mode :character Median : -1.00 Median :2014 Median : 6.000
## Mean : 12.47 Mean :2014 Mean : 5.639
## 3rd Qu.: 11.00 3rd Qu.:2014 3rd Qu.: 8.000
## Max. :1498.00 Max. :2014 Max. :10.000
##
## day arr_delay origin dest
## Min. : 1.00 Min. :-112.000 Length:253316 Length:253316
## 1st Qu.: 8.00 1st Qu.: -15.000 Class :character Class :character
## Median :16.00 Median : -4.000 Mode :character Mode :character
## Mean :15.89 Mean : 8.147
## 3rd Qu.:23.00 3rd Qu.: 15.000
## Max. :31.00 Max. :1494.000
##
## air_time distance hour sum_delay
## Min. : 20.0 Min. : 80.0 Min. : 0.00 Min. :-224.00
## 1st Qu.: 86.0 1st Qu.: 529.0 1st Qu.: 9.00 1st Qu.: -19.00
## Median :134.0 Median : 762.0 Median :13.00 Median : -5.00
## Mean :156.7 Mean : 950.4 Mean :13.06 Mean : 20.61
## 3rd Qu.:199.0 3rd Qu.:1096.0 3rd Qu.:17.00 3rd Qu.: 23.00
## Max. :706.0 Max. :4963.0 Max. :24.00 Max. :2992.00
## NA's :81483
## carrierm_mean_arr carrierm_mean_dep lkp.count
## Min. :-22.403 Min. :-4.500 Min. : 85
## 1st Qu.: 2.676 1st Qu.: 7.815 1st Qu.:3449
## Median : 6.404 Median :11.354 Median :5877
## Mean : 8.147 Mean :12.465 Mean :4654
## 3rd Qu.: 11.554 3rd Qu.:17.564 3rd Qu.:6058
## Max. : 86.182 Max. :52.864 Max. :6058
## NA's :229797
Cardinality
We can check the uniqueness of data by using uniqueN function and apply it on every column. Object .SD in the query below corresponds to Subset of the Data.table:
DT[, lapply(.SD, uniqueN)]
## carrier dep_delay year month day arr_delay origin dest air_time
## 1: 14 570 1 10 31 616 3 109 509
## distance hour sum_delay carrierm_mean_arr carrierm_mean_dep lkp.count
## 1: 152 25 1021 134 134 11
NA Ratio
To calculate the ratio of unknown values (NA in R, and NULL in SQL) for each column, we provide the desired function to apply on every column.
DT[, lapply(.SD, function(x) sum(is.na(x))/.N)]
## carrier dep_delay year month day arr_delay origin dest air_time
## 1: 0 0 0 0 0 0 0 0 0
## distance hour sum_delay carrierm_mean_arr carrierm_mean_dep lkp.count
## 1: 0.3216654 0 0 0 0 0.9071555
Exporting Data
Fast export tabular data to CSV format is also provided by the data.table package.
tmp.csv <- tempfile(fileext=".csv")
fwrite(DT, tmp.csv)
# preview exported data
cat(system(paste("head -3",tmp.csv), intern=TRUE), sep="\n")
## carrier,dep_delay,year,month,day,arr_delay,origin,dest,air_time,distance,hour,sum_delay,carrierm_mean_arr,carrierm_mean_dep,lkp.count
## AA,1498,2014,10,4,1494,EWR,DFW,200,1372,7,2992,5.54195933456561,7.59149722735674,5877
## AA,1241,2014,4,15,1223,JFK,BOS,39,,13,2464,1.90332441727168,3.98700802445548,1173
At the time of writing this, the fwrite function hasn’t yet been published to the CRAN repository. To use it we need to install data.table development version, otherwise we can use base R write.csv function, but don’t expect it to be fast.
Resources
There are plenty of resources available. Besides the manuals available for each function, there are also package vignettes, which are tutorials focused around the particular subject. Those can be found on the Getting started page. Additionally, the Presentations page lists more than 30 materials (slides, video, etc.) from data.table presentations around the globe. Also, the community support has grown over the years, recently reaching the 4000-th question on Stack Overflow data.table tag, still having a high ratio (91.9%) of answered questions. The below plot presents the number of data.table tagged questions on Stack Overflow over time.
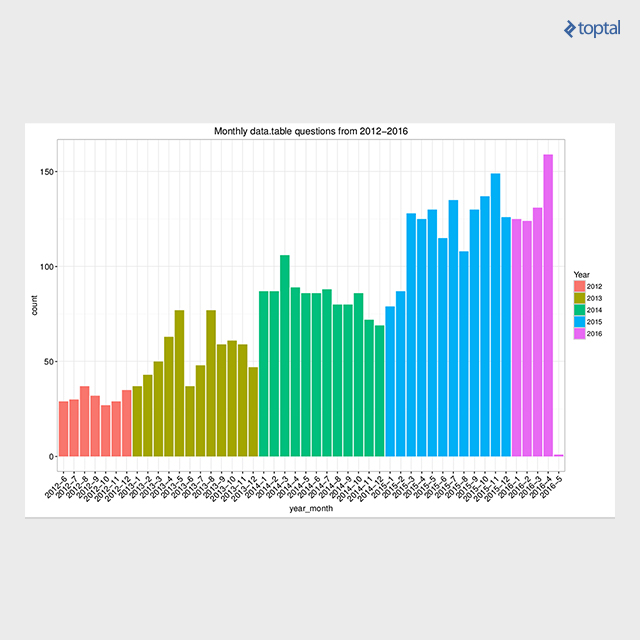
Summary
This article provides chosen examples for efficient tabular data transformation in R using the data.table package. The actual figures on performance can be examined by looking for reproducible benchmarks. I published a summarized blog post about data.table solutions for the top 50 rated StackOverflow questions for the R language called Solve common R problems efficiently with data.table, where you can find a lot of figures and reproducible code. The package data.table uses native implementation of fast radix ordering for its grouping operations, and binary search for fast subsets/joins. This radix ordering has been incorporated into base R from version 3.3.0. Additionally, the algorithm was recently implemented into H2O machine learning platform and parallelized over H2O cluster, enabling efficient big joins on 10B x 10B rows.

